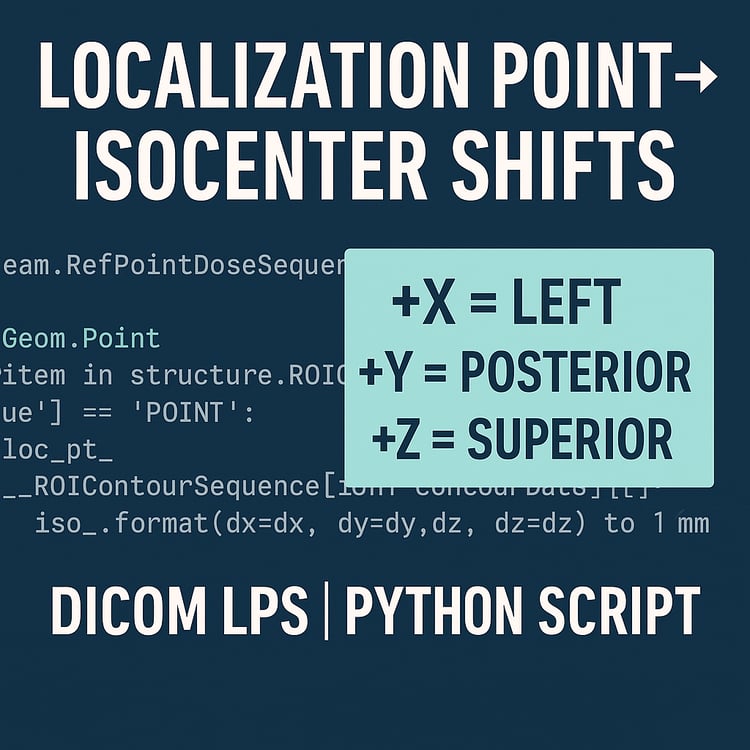
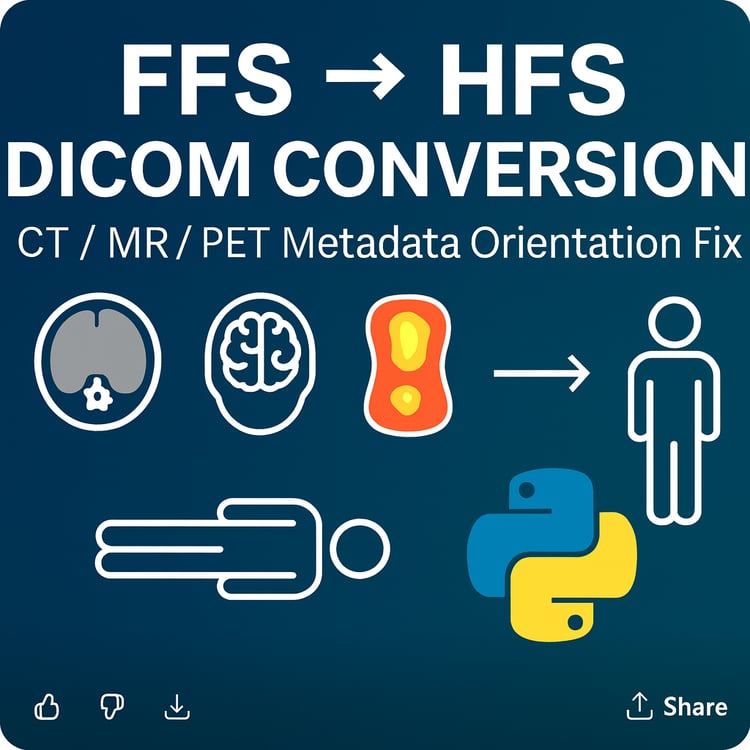
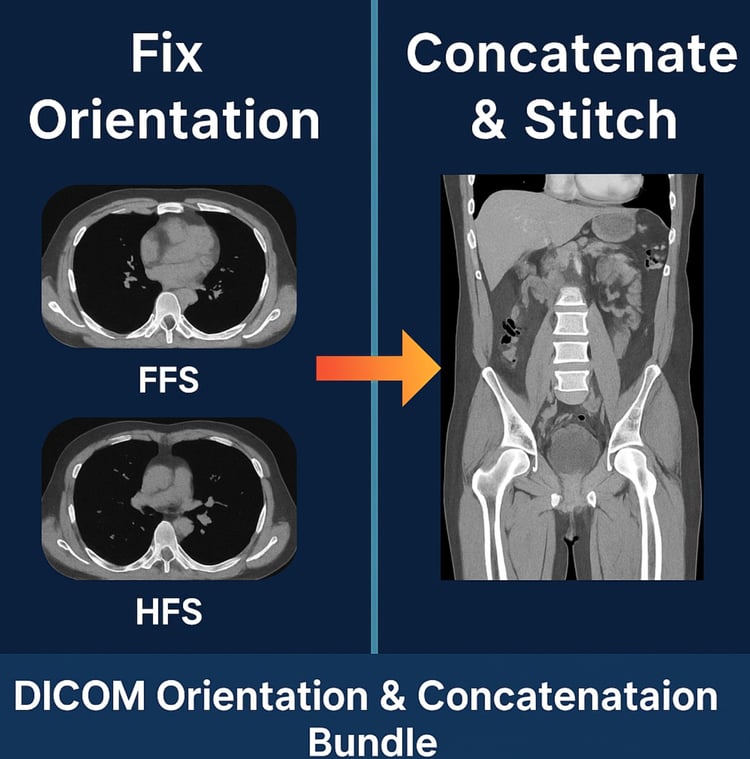
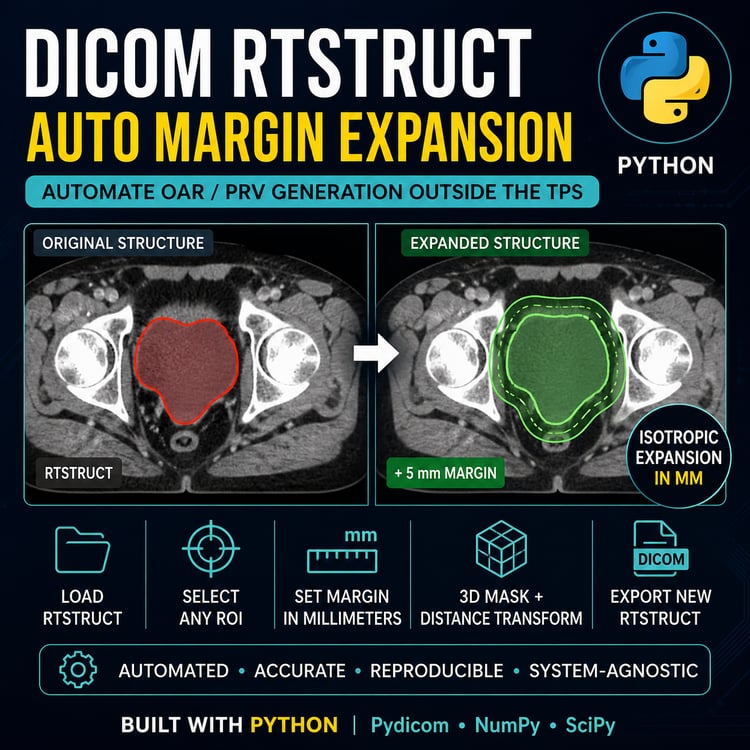
EQD2 Dose Converter (Python Script) — Convert RTDOSE to EQD2 Directly from DICOM
RTDOSE → EQD2 Conversion Tool (Global & OAR-Specific α/β)
Convert RTDOSE → EQD2 in seconds with this Python-based, fully DICOM-native tool.
This script provides a TPS-independent workflow for computing biologically equivalent dose (EQD2) distributions directly from DICOM RTDOSE files — with support for both global and organ-specific α/β values.
Whether you’re validating radiobiological models, performing independent dose checks, or building automation workflows, this tool generates clean, import-ready EQD2 RTDOSE files that can be viewed in any DICOM-compliant system.
✨ Key Features
✔️ Fully DICOM-native implementation (pydicom — no TPS required)
✔️ Voxel-by-voxel RTDOSE → EQD2 conversion using the linear-quadratic model
✔️ Global EQD2 calculation using a single α/β value applied to the entire dose grid
✔️ OAR-specific α/β overrides
• Apply different α/β values inside selected organs at risk
• Retain the global α/β everywhere else
✔️ Generates multiple EQD2 dose series for direct comparison:
• Original physical dose
• EQD2 (global α/β only)
• EQD2 (with OAR-specific α/β overrides)
✔️ Automatic DoseGridScaling calculation for safe DICOM export
✔️ Preserves FrameOfReferenceUID for correct CT/dose alignment
✔️ Generates new DICOM UIDs for clean import into any system
✔️ Removes ReferencedRTPlanSequence so the dose is treated as an evaluation dose
✔️ Produces RTDOSE files compatible with all DICOM-compliant TPS, PACS, and viewers
✔️ Clean, well-commented Python code — easy to customize for research or clinical workflows
🎯 Perfect For
• Medical physicists
• Radiobiology researchers
• DICOM developers
• Automation and scripting engineers
• Anyone needing biological dose visualization outside the TPS
🛠️💡 Need customization? Contact me to adapt this tool to your workflow.